Abstract
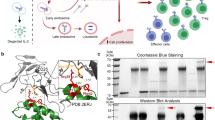
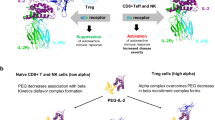
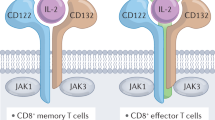
The immunostimulatory cytokine interleukin-2 (IL-2) is a growth factor for a wide range of leukocytes, including T cells and natural killer (NK) cells1,2,3. Considerable effort has been invested in using IL-2 as a therapeutic agent for a variety of immune disorders ranging from AIDS to cancer. However, adverse effects have limited its use in the clinic. On activated T cells, IL-2 signals through a quaternary ‘high affinity’ receptor complex consisting of IL-2, IL-2Rα (termed CD25), IL-2Rβ and IL-2Rγ4,5,6,7,8. Naive T cells express only a low density of IL-2Rβ and IL-2Rγ, and are therefore relatively insensitive to IL-2, but acquire sensitivity after CD25 expression, which captures the cytokine and presents it to IL-2Rβ and IL-2Rγ. Here, using in vitro evolution, we eliminated the functional requirement of IL-2 for CD25 expression by engineering an IL-2 ‘superkine’ (also called super-2) with increased binding affinity for IL-2Rβ. Crystal structures of the IL-2 superkine in free and receptor-bound forms showed that the evolved mutations are principally in the core of the cytokine, and molecular dynamics simulations indicated that the evolved mutations stabilized IL-2, reducing the flexibility of a helix in the IL-2Rβ binding site, into an optimized receptor-binding conformation resembling that when bound to CD25. The evolved mutations in the IL-2 superkine recapitulated the functional role of CD25 by eliciting potent phosphorylation of STAT5 and vigorous proliferation of T cells irrespective of CD25 expression. Compared to IL-2, the IL-2 superkine induced superior expansion of cytotoxic T cells, leading to improved antitumour responses in vivo, and elicited proportionally less expansion of T regulatory cells and reduced pulmonary oedema. Collectively, we show that in vitro evolution has mimicked the functional role of CD25 in enhancing IL-2 potency and regulating target cell specificity, which has implications for immunotherapy.
This is a preview of subscription content, access via your institution
Access options
Access to this article via Institution of Civil Engineers Library is not available.
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
16 April 2012
In the legend to Supplementary Movie 2 'wild-type IL-2' was changed to 'D10 IL-2 superkine'.
References
Rochman, Y., Spolski, R. & Leonard, W. J. New insights into the regulation of T cells by γc family cytokines. Nature Rev. Immunol. 9, 480–490 (2009)
Smith, K. A. Interleukin-2: inception, impact, and implications. Science 240, 1169–1176 (1988)
Waldmann, T. A. The biology of interleukin-2 and interleukin-15: implications for cancer therapy and vaccine design. Nature Rev. Immunol. 6, 595–601 (2006)
Cosman, D. et al. Cloning, sequence and expression of human interleukin-2 receptor. Nature 312, 768–771 (1984)
Leonard, W. J. et al. Molecular cloning and expression of cDNAs for the human interleukin-2 receptor. Nature 311, 626–631 (1984)
Nikaido, T. et al. Molecular cloning of cDNA encoding human interleukin-2 receptor. Nature 311, 631–635 (1984)
Hatakeyama, M. et al. Interleukin-2 receptor beta chain gene: generation of three receptor forms by cloned human alpha and beta chain cDNA’s. Science 244, 551–556 (1989)
Takeshita, T. et al. Cloning of the gamma chain of the human IL-2 receptor. Science 257, 379–382 (1992)
Boder, E. T. & Wittrup, K. D. Yeast surface display for screening combinatorial polypeptide libraries. Nature Biotechnol. 15, 553–557 (1997)
Chao, G. et al. Isolating and engineering human antibodies using yeast surface display. Nature Protocols 1, 755–768 (2006)
Wang, X., Rickert, M. & Garcia, K. C. Structure of the quaternary complex of interleukin-2 with its α, β, and γc receptors. Science 310, 1159–1163 (2005)
Mott, H. R. et al. The solution structure of the F42A mutant of human interleukin 2. J. Mol. Biol. 247, 979–994 (1995)
Thanos, C. D., DeLano, W. L. & Wells, J. A. Hot-spot mimicry of a cytokine receptor by a small molecule. Proc. Natl Acad. Sci. USA 103, 15422–15427 (2006)
Boyman, O., Kovar, M., Rubinstein, M. P., Surh, C. D. & Sprent, J. Selective stimulation of T cell subsets with antibody-cytokine immune complexes. Science 311, 1924–1927 (2006)
Krieg, C., Létourneau, S., Pantaleo, G. & Boyman, O. Improved IL-2 immunotherapy by selective stimulation of IL-2 receptors on lymphocytes and endothelial cells. Proc. Natl Acad. Sci. USA 107, 11906–11911 (2010)
Létourneau, S. et al. IL-2/anti-IL-2 antibody complexes show strong biological activity by avoiding interaction with IL-2 receptor α subunit CD25. Proc. Natl Acad. Sci. USA 107, 2171–2176 (2010)
Rosenberg, S. A., Mule, J. J., Spiess, P. J., Reichert, C. M. & Schwarz, S. L. Regression of established pulmonary metastases and subcutaneous tumor mediated by the systemic administration of high-dose recombinant interleukin 2. J. Exp. Med. 161, 1169–1188 (1985)
Rao, B. M., Driver, I., Lauffenburger, D. A. & Wittrup, K. D. Interleukin 2 (IL-2) variants engineered for increased IL-2 receptor α-subunit affinity exhibit increased potency arising from a cell surface ligand reservoir effect. Mol. Pharmacol. 66, 864–869 (2004)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Rickert, M., Boulanger, M. J., Goriatcheva, N. & Garcia, K. C. Compensatory energetic mechanisms mediating the assembly of signaling complexes between interleukin-2 and its α, β, and γc receptors. J. Mol. Biol. 339, 1115–1128 (2004)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007)
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007)
DeLano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific, 2002)
Eswar, N. et al. Comparative Protein Structure Modeling With MODELLER. Current Protocols in Bioinformatics. Vol. 15 5.6.1-5.6.30 (Wiley, 2006)
Bazan, J. F. Unraveling the structure of IL-2. Science 257, 410–413 (1992)
Arkin, M. R. et al. Binding of small molecules to an adaptive protein–protein interface. Proc. Natl Acad. Sci. USA 100, 1603–1608 (2003)
Rickert, M., Wang, X., Boulanger, M. J., Goriatcheva, N. & Garcia, K. C. The structure of interleukin-2 complexed with its alpha receptor. Science 308, 1477–1480 (2005)
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008)
Duan, Y. et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 24, 1999–2012 (2003)
Hess, B. P-LINCS: a parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 4, 116–122 (2008)
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Hoover, W. G. Canonical dynamics: equilibrium phase-space distributions. Phys. Rev. A 31, 1695–1697 (1985)
Berendsen, H. J. C., Postma, P. M., van Gunsteren, W. F., DiNola, A. & Haak, J. R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984)
Bowman, G. R., Huang, X. & Pande, V. S. Using generalized ensemble simulations and Markov state models to identify conformational states. Methods 49, 197–201 (2009)
Bowman, G. R., Beauchamp, K. A., Boxer, G. & Pande, V. S. Progress and challenges in the automated construction of Markov state models for full protein systems. J. Chem. Phys. 131, 124101 (2009)
Bowman, G. R., Huang, X. & Pande, V. S. Network models for molecular kinetics and their initial applications to human health. Cell Res. 20, 622–630 (2010)
Noé, F. & Fischer, S. Transition networks for modeling the kinetics of conformational change in macromolecules. Curr. Opin. Struct. Biol. 18, 154–162 (2008)
Acknowledgements
The authors gratefully acknowledge W. Leonard, R. Levy and R. Schwendener for reagents and discussion. This work was supported by NIH-RO1AI51321 (to K.C.G.), PP00P3-128421 from the Swiss National Science Foundation and KFS-02672-08-2010 from the Swiss Cancer League (both to O.B.), NIH R01-GM062868 (to V.S.P.), MRI-R2 (this award is funded under the American Recovery and Reinvestment Act of 2009 (Public Law 111-5)) (to V.S.P.), NIH-AR050942 (to J.T.L.), NIH U01 DK078123 (to C.G.F.), and NIH U19 AI 082719 (to C.G.F.). A.M.R. was supported by the Stanford Medical Scientist Training Program (NIH-GM07365). K.C.G. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
A.M.L. performed in vitro evolution and contributed to preparation of the manuscript. D.L.B. produced recombinant proteins, determined crystal structures, and carried out surface plasmon resonance analysis. A.M.R. carried out cellular and signalling assays, biophysical measurements and contributed to preparation of the manuscript. C.K. carried out in vivo experiments, analysed data and contributed to preparation of the manuscript; M.E.R. carried out in vivo experiments in mice. I.M. analysed cell-signalling data. G.R.B., P.N. and V.S.P. carried out and analysed molecular dynamics simulations. J.T.L., L.S. and C.G.F. performed and analysed T-cell signalling experiments. O.B. designed and supervised in vivo experiments, analysed data and contributed to preparation of the manuscript. K.C.G. conceived of the project, analysed data, supervised execution of the project, and prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
K.C.G., A.M.L. and A.M.R. declare competing financial interests due to submission of a pending patent application describing the IL-2 superkine. O.B. declares competing financial interests due to being a shareholder of Nascent Biologics Inc.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, Supplementary Figures 1-12 and the legend for Supplementary Movies 1-2. (PDF 6476 kb)
Supplementary Movie 1
In this movie we see atomistic molecular dynamics simulations of wild-type IL-2 (see Supplementary Information file for movie legend). (MOV 956 kb)
Supplementary Movie 2
In this movie we see atomistic molecular dynamics simulations of D10 IL-2 superkine (see Supplementary Information file for movie legend). (MOV 499 kb)
Rights and permissions
About this article
Cite this article
Levin, A., Bates, D., Ring, A. et al. Exploiting a natural conformational switch to engineer an interleukin-2 ‘superkine’. Nature 484, 529–533 (2012). https://doi.org/10.1038/nature10975
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10975
This article is cited by
-
Molecular Engineering of Interleukin-2 for Enhanced Therapeutic Activity in Autoimmune Diseases
BioDrugs (2024)
-
NK cells as powerful therapeutic tool in cancer immunotherapy
Cellular Oncology (2024)
-
New insights into the stemness of adoptively transferred T cells by γc family cytokines
Cell Communication and Signaling (2023)
-
The decreased of peripheral blood natural killer cell is associated with serum IL-2 level in the renal tubular acidosis in patients with primary sjogren’s syndrome
BMC Immunology (2023)
-
Emerging principles of cytokine pharmacology and therapeutics
Nature Reviews Drug Discovery (2023)